Improved fiber tractography algorithm for neurosurgical planning
Andrey Zhylka defended his thesis at the Department of Biomedical Engineering on September 8th.

Accurate fiber bundle reconstruction is required for neurosurgery planning to minimize the risk of damaging healthy nerve bundles. When choosing a fiber tractography algorithm for the reconstruction, there is currently a trade-off between precision and the extent of reconstruction. For his PhD research, Andrey Zhylka proposed a multi-level fiber tractography algorithm that reconstructs pathways by adding the capability of including branching pathways. Given evaluation on healthy subjects and clinical patients, it was shown to be robust, produce a feasible reconstruction extent, and preserve fiber bundle topography, therefore lowering the risk of introducing unnecessary brain damage.
The human brain consists of an enormous number of axonal fibers organized in bundles. Each bundle is responsible for one or several functions of the human body. The integrity of such fibers is essential for the brain to be functioning. Brain tumors can suppress the activity of fiber bundles or even cause their disruption. Neurosurgery is the typical treatment for brain tumors to either fully or partially recover the well-being of the patient.
However, neurosurgery can damage unaffected areas of the brain as surgeons aim to maximize tumor resection. To minimize the removal of healthy tissue, surgeons need tools that can provide them with information on the organization of the fiber bundles and whether or not they are affected by the tumor.
Imaging
Diffusion MRI is an imaging technique that can provide such information. Using nerve fiber reconstruction algorithms (called fiber tractography algorithms), it is possible to obtain a 3D reconstruction of the fiber bundles and their properties. Ideally, neurosurgeons could integrate this information into the surgical planning to adapt the extent of resection and optimize the trajectory of the intervention.
However, currently, when choosing a tractography algorithm one faces a trade-off between the precision and the extent of reconstruction. Hence, in this work, we aimed to develop a method that improves the completeness of fiber bundle reconstructions while alleviating the limitations of the existing advanced algorithms.
Improving reconstruction
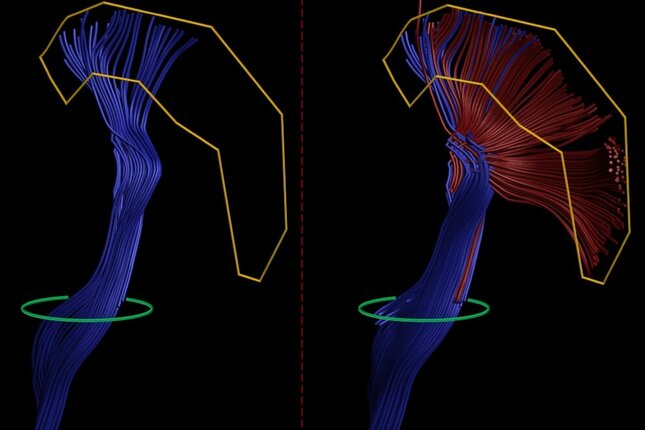
The multi-level tractography framework presented in this work gradually improves fiber bundle reconstruction by letting already reconstructed fibers branch. At the same time, to control the accuracy the reconstructed fibers are filtered based on neuroanatomical knowledge about the target fiber bundle, namely which brain regions it connects to.
The framework was evaluated on data from healthy subjects and data from patients with brain tumors. It produced adequate reconstructions that are more complete compared to those obtained with the algorithm routinely used in the clinic.
Given the evaluation results, the multi-level tractography can provide more complete reconstructions that allow us to estimate a safer resection margin or to point out areas that would require additional attention during surgery. Hence, the risk of introducing unnecessary brain damage becomes lower.
Title of PhD thesis: “Diffusion MRI tractography branched out”
Supervisors: Josien Pluim and Marcel Breeuwer